I started a climate modeling project assuming I’d be dealing with “large” datasets. Then I saw the actual size: 2 terabytes. I wrote a straightforward NumPy script, hit-run, and grabbed a coffee. Bad idea. When I came back, my machine had frozen. I restarted and tried a smaller slice. Same crash. My usual workflow wasn’t going to work. After some trial and error, I eventually landed on Zarr, a Python library for chunked array storage. It let me process that entire 2TB dataset on my laptop without any crashes. Here’s what I learned:
When “big data” actually means big trouble
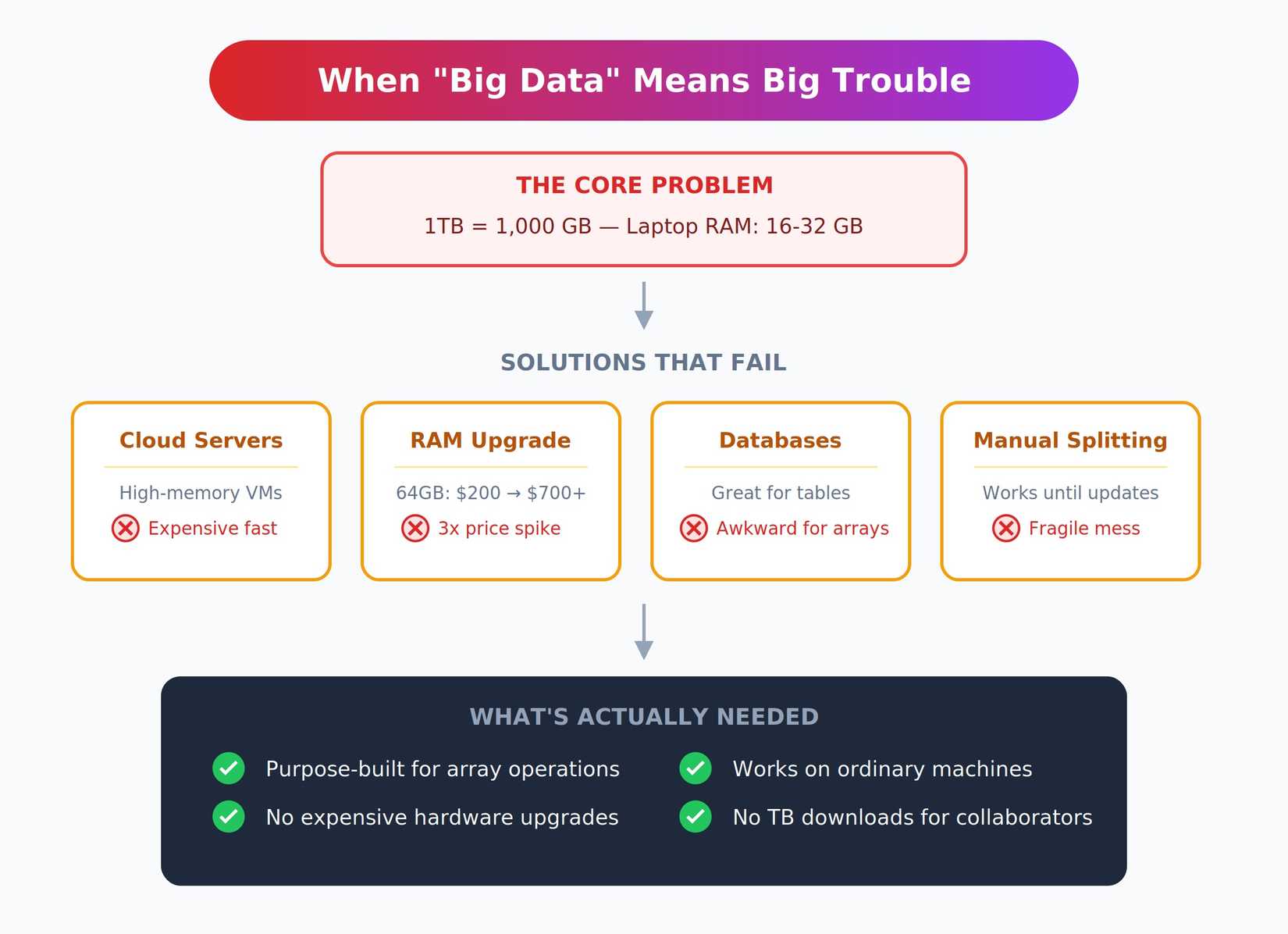
Working with terabytes isn’t just “more data.” It’s an entirely different class of problem that breaks all your normal assumptions.
Most laptops have 16 to 32 GB of RAM. A single terabyte is roughly 1,000 GB. You simply cannot load that into memory, no matter how clever your code is. Even if you could somehow squeeze it in, sequential reads at this scale are painfully slow.
The usual solutions all have serious drawbacks. You could rent a high-memory cloud server with hundreds of gigabytes of RAM, but that gets expensive fast, and personally upgrading your local machine’s RAM is no longer a cost-effective alternative. RAM prices have roughly tripled since mid-2025 due to AI data centers consuming most of the supply.
That 64GB upgrade that cost $200 last spring now runs over $700. Databases work great for tabular data, but feel awkward for multidimensional arrays. Manually splitting files into manageable pieces works until you need to update something, then it becomes a fragile mess.
I needed something purpose-built for array operations that didn’t require expensive hardware upgrades. More importantly, I needed collaborators to access results on their ordinary machines without downloading terabytes of files first.
Meet Zarr (finally, something designed for this)
Zarr is a Python library designed for large, chunked array storage. The core idea is beautifully simple: break arrays into independently stored chunks. Each chunk can be compressed on its own and read separately from the rest. You interact with a Zarr array almost exactly like a NumPy array, with familiar slicing and indexing syntax. But under the hood, Zarr only loads the chunks you actually need into memory.
The library supports local disks, network drives, and cloud backends like S3, Google Cloud Storage, or Azure Blob. This made it possible to process cloud-hosted data without downloading everything first. For a 2TB dataset, that capability alone was game-changing.
Zarr is open source, actively maintained by the scientific Python community, and plays well with the existing ecosystem. The API feels familiar, not like learning an entirely new system. If you know NumPy, you’re already most of the way there.
Stop crashing your Python scripts: How Zarr handles massive arrays
Tired of out-of-memory errors derailing your data analysis? There’s a better way to handle huge arrays in Python.
Putting it to the test on real data
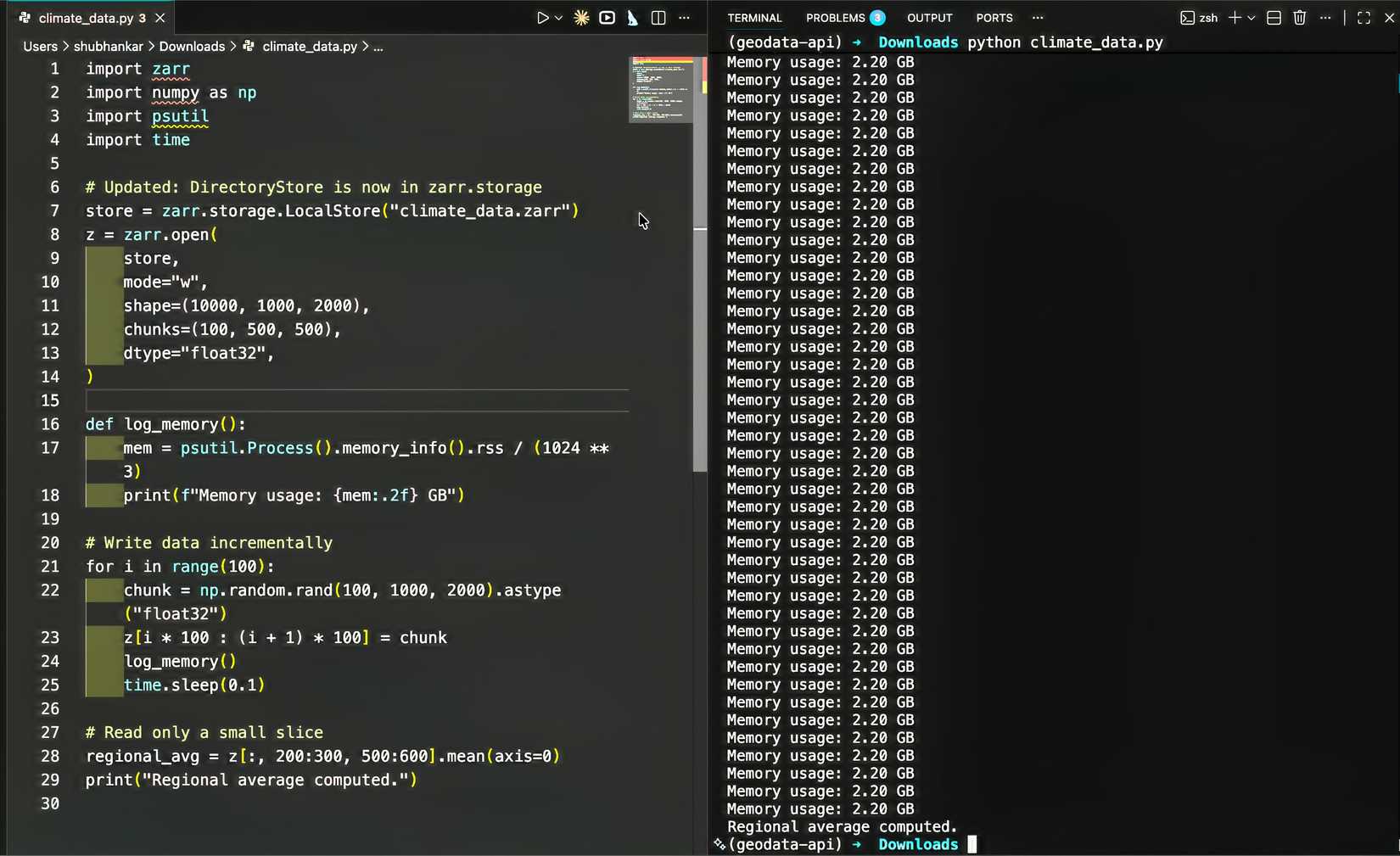
My test dataset was about 2TB of climate simulation output spanning thousands of time steps. The goal was straightforward: calculate regional averages across the entire time series. I set up a Zarr array with carefully chosen chunk sizes. This took some experimentation. Chunks that are too small add overhead from managing too many files. Chunks that are too large defeat the purpose by forcing you to load gigabytes into memory at once.
I eventually settled on chunks that matched my access patterns, slicing by time and geographic region. The actual code looked remarkably similar to my original NumPy script:
import zarr
import numpy as np
# Create a chunked Zarr array
store = zarr.DirectoryStore('climate_data.zarr')
z = zarr.open(store, mode='w', shape=(10000, 1000, 2000),
chunks=(100, 500, 500), dtype='float32')
# Write data in chunks
for i in range(100):
chunk_data = process_raw_data(i)
z[i*100:(i+1)*100] = chunk_data
# Read and process specific slices
regional_avg = z[:, 200:300, 500:600].mean(axis=0)The first full run was completed successfully in a few hours. My laptop’s memory usage stayed around 4 GB the entire time. No crashes, no freezes, just steady progress through the dataset.

Learn Python Basics: Let’s Build A Simple Expense Tracker
Take charge of your finances while learning to code in Python.
The Zarr features that actually mattered
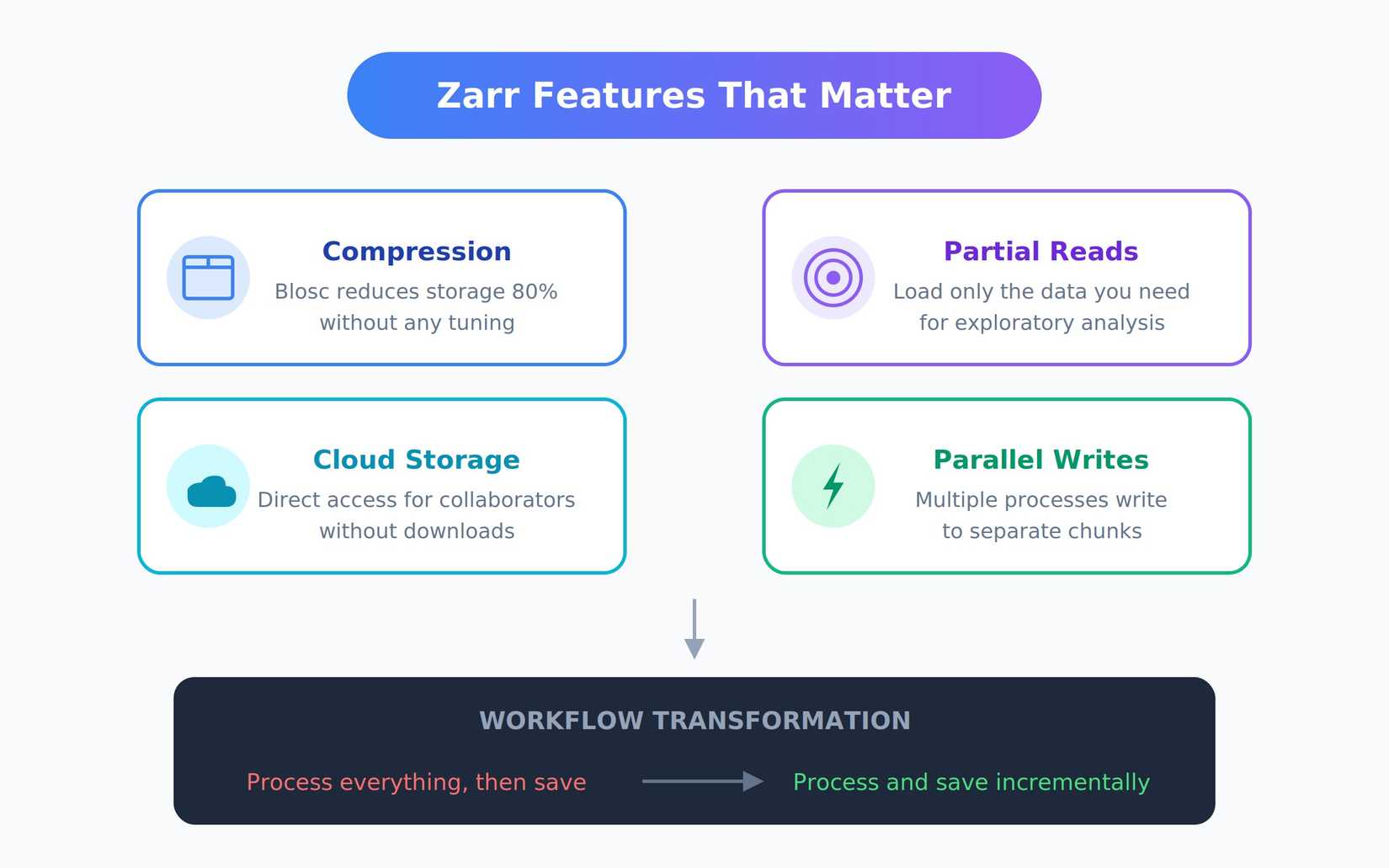
Compression was surprisingly effective without any tuning. Zarr uses Blosc by default, which reduced my 2TB dataset to under 400GB. For scientific data with patterns and repetition, compression ratios like this are common. Partial reads made exploratory analysis possible. I could load just January data, or just a specific region, without touching the rest. What was previously impossible became routine.
Cloud storage support eliminates local disk bottlenecks. I moved the final array to Google Cloud Storage. Collaborators could open it directly without downloads, and I stopped worrying about backup strategies for multi-terabyte files.
Parallel writes let me split processing across multiple scripts or machines. Different processes could write to separate chunks simultaneously without conflicts. This turned a multi-day sequential job into something I could finish in hours. Eventually, my workflow changed. Instead of “process everything, then save,” I shifted to “process and save incrementally.” Zarr works just as well for building arrays piece by piece as it does for reading them back.
Where Zarr makes you work a little harder

Despite its many advantages, Zarr isn’t without its tradeoffs. You still need to think carefully about chunk size and access patterns. Poor choices can make small workloads slower than plain NumPy. Heavy random access across many chunks is expensive. If you’re constantly jumping around unpredictably, you’ll pay the cost of loading different chunks repeatedly. Some problems require rethinking your approach.
There’s a real learning curve around performance tuning.
Documentation is solid but spread across Zarr, Xarray for labeled arrays, and Dask for parallel computing. Figuring out which tool handles what takes time. In practice, Zarr often works best paired with Dask for parallel operations or Xarray for dimension labels. The overlap between tools can be confusing for newcomers. For my workload, which involved mostly sequential access and regional aggregations, the trade-offs were absolutely worth it. But if your problem fits comfortably in RAM already, stick with NumPy.
Zoom out: Why this changed my default workflow
Zarr isn’t the only solution. HDF5 has been around longer. TileDB targets similar problems with different design choices. NetCDF4 remains standard in climate science. What made Zarr stand out was how naturally it fit into the Python ecosystem. Xarray adds dimension labels and coordinate systems, and Dask adds parallelism and distributed computing. The pieces connect smoothly.
Working at a terabyte scale no longer felt unusual, and I default to Zarr once datasets exceed a few gigabytes, even if they technically fit in RAM. The convenience of compression and cloud storage alone makes it worthwhile. The scientific Python stack has matured into a serious big data platform. You don’t need Hadoop or Spark for array workloads anymore.
Zarr saved the project and probably saved my sanity. What felt impossible on my laptop became manageable. If you work with large arrays and keep hitting memory limits, Zarr is worth exploring. The learning curve pays for itself quickly, especially considering current RAM prices make hardware upgrades prohibitively expensive.
Start with the official Zarr documentation and experiment with chunking strategies on smaller datasets first. You can start small and scale up as needed without rewriting everything. Once comfortable with the basics, the broader ecosystem of Xarray and Dask opens up even more possibilities.






